I’m really excited to have been recently awarded an Alfred P. Sloan Foundation Microbiology of the Built Environment postdoctoral fellowship. I was asked to explain a little about my plans for the project I’ll be working on for the next two years now that I officially have my PhD.
As a graduate student I investigated the factors that affect the microbial communities of breast-fed infants. Babies go from being near-sterile in the womb to all of a sudden encountering the full microbial diversity of their new ex-utero environment, yet retain a gut microbiota distinct from that of adults. Our microbiota likely coevolved with us, and influences our own physiology in a number of ways1. As a PhD student in the lab of David Mills at UC Davis, I studied the microbiota of several infant cohorts from different areas of the world. Out of curiosity, I compared the data I worked on (from infants in Bangladesh2, the USA3, and a few as-yet unpublished datasets) with several other published datasets. I noticed a trend showing differences between the gut microbiomes of infants in Western, Educated, Industrialized, Rich, and Democratic (“WEIRD”) nations and nations in the Global South4. Most notably, healthy breast-fed infants in WEIRD populations had higher amounts of Bifidobacterium than infants in the Global South — a phenomenon that had been noted before5.
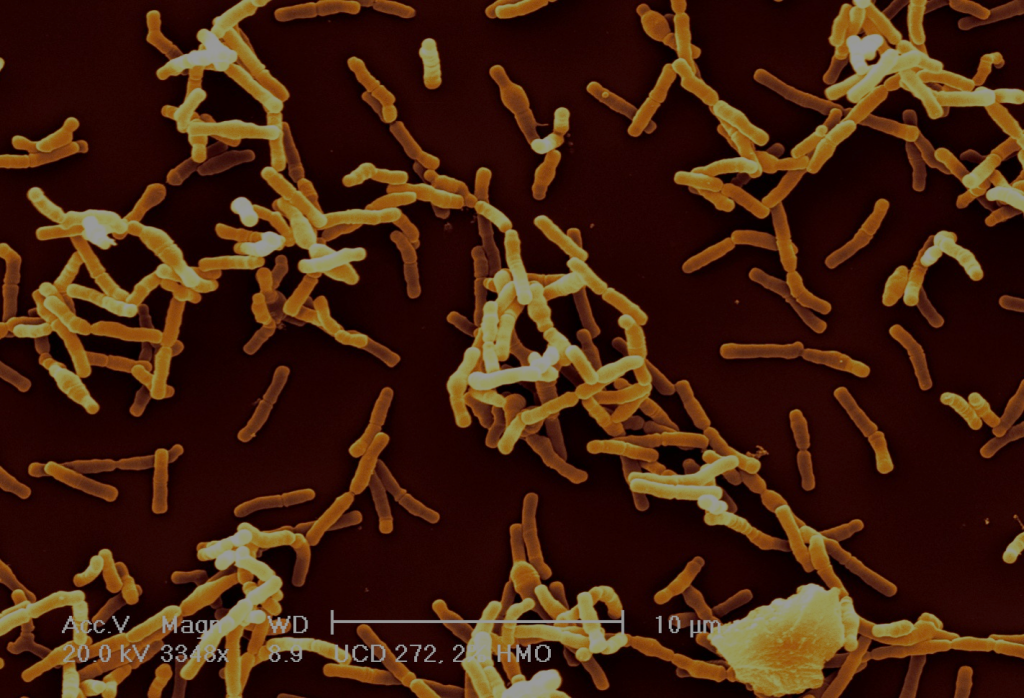
Bifidobacteria are gram-positive anaerophilic bacteria, and are interesting for a number of reasons. First off, they are very common in infants, much more so than in adults. Studies using marker gene sequencing techniques show that the gut of a healthy breast-fed infant is often dominated by bifidobacteria 6—9. This is likely because bifidobacteria are one of the few types of bacteria that can fully consume complex human milk glycans (HMGs) found in breast milk which are indigestible to the infant10—15. The structural complexity of HMGs serves as a barrier for other microbes to compete with bifidobacteria for these sugars in the infant gut. Bifidobacteria benefit from colonizing infants by getting access to these sugars — their “favorite food.”
But what is in it for the infants and mothers? Why bother to provide these sugars to the bacteria? There are some ideas as to why. Infants with bifidobacteria-dominated gastrointestinal tracts have a higher resistance to colonization by pathogens, improved responses to some vaccines, and better gut barrier function 2,15—17. Bifidobacteria aid the appropriate development of the infant’s innate and acquired immune systems, simultaneously enhancing surveillance yet reducing inflammation 17—20. Infant-type bifidobacteria present during weaning may guide the immune system towards tolerance during the introduction of new foods and their associated antigens 21—24. I think levels of bifidobacteria might be suitable as a biomarker for a desirable infant gut microbiome, even though the ecological dynamics at play within the infant microbiota and their relationship to health outcomes are complex and not fully understood. For that reason, the observed differences in bifidobacterial levels across the globe might have some important public health implications.
Additional studies are needed to confirm this trend of lower amounts of bifidobacteria in infants in the WEIRD world, but if it holds true, identifying the cause(s) will be important for understanding this phenomenon. I see two broad possibilities for what the cause(s) might be: either the gut environments of infants in the WEIRD nations are differentially selective (against bifidobacteria), or there are higher barriers to bifidobacteria getting into infants in the WEIRD world (bifidobacteria are in fact, not “everywhere25”). My project focuses on the second possibility. The initial source of microbes in infants is often thought to be a combination of the maternal vaginal microbiota 26,27, the mother’s intestinal microbiota 28, and other environmental factors including caretaker skin 29. Breast milk also contains microbes, but their origin and potential impact on colonization is unclear 30—32. However, I’d like to test an alternative hypothesis — that the most likely place from which an infant might acquire infant-adapted bifidobacteria is… another infant. Infants in WEIRD nations might have lower propinquity to other infants than babies in the Global South, and thus have challenges in acquiring bifidobacteria from them. This low infant/infant overlap lifestyle may differ from the conditions experienced over the course of evolutionary history, and it may be (in part) driving the lower levels of bifidobacterial colonization I’ve observed in WEIRD places.
But how can this hypothesis be tested? The potential of the built environment as venue for cross-inoculation between infants has not been studied, despite hints of shared microbes in the built environment and infant gut ecosystems33. Could infants be leaving a microbial trace of their presence on the environmental features with which they interact? Barring outside intervention, infants are indiscriminate in their spread of fecal microbes to individuals and objects with which they come into contact (as any parent that has experienced a major diaper blow-out can testify). My plan is to examine specific built environments to investigate potential microbial signatures left by infants. I will also test the microbiomes of infants in the U.S.A. and compare them to those of the Himba, a population of traditionally-living pastoralists. There are two main questions I am asking in this study:
Question 1: Do breast-fed infants leave viable bifidobacteria in their surrounding built environment which are not found in non-infant-associated built environments?
To investigate the nature of the infant-associated built environment microbiome, I will study the microbiota of daycare centers. I will swab select built environment features (play tables, toys, floors, etc.) and analyze the microbes captured through sequencing and culture-based methods (to test viability). I will collect metadata about potential microbiota-influencing building conditions at these locations (e.g. usage, cleaning procedures, relative humidity, and temperature) using building sensors developed at UC Davis (thanks to Nic Madrid and Prof. Andre Knoesen). As a positive control, I will also study diaper-changing tables (somewhere where the presence of infant fecal microbes might be expected) whose usage will also be monitored with sensors. As a negative control, I will also be studying a series of 40 dedicated lactation sites on UC Davis campus which are used by breastfeeding mothers to express breast milk. A special thanks to the UC Davis breastfeeding support program for giving me and my team of undergraduates permission to swab the rooms. As infants are not typically present in these rooms, these spaces serve as a negative control where direct deposition of microbes by infants is not a factor. However, as the rooms are used principally by mothers who come into frequent contact with their own breast-fed infants, studying the lactation sites would allow for detection of potential indirect transfer of microbes between infants via the mothers.
Question 2: Are levels of bifidobacteria in the infant gut be positively associated with exposure to multi-infant built environments?
I plan to address this question in two ways. First, by comparing infants with different exposures within the USA. Approximately 80 infants are currently enrolled in the UC Davis Lactation Study. Fecal samples from the infants were collected over the first year of life, and intensive metadata was collected about breastfeeding, C-section births, antibiotic supplementation by the mother and infant, maternal use of personal products containing antimicrobial agents, maternal demographics, and numerous other factors. However, information about daycare enrollment, social interactions (e.g. “mommy-and-me” groups), shared nannies, and other social factors was not collected. Together with the leaders of the lactation study, I have sent out a survey to the mothers asking for this additional information so that I can compare it to data on the infants’ microbiomes.
The second way I want to ask this question is by comparing the US infants as a whole to infants in a very different setting, the Himba of Namibia. The Himba are traditionally-living pastoralists that live a very different lifestyle from people in WEIRD countries. I will explore the impact of various social factors on the infant microbiome through the sampling of infant feces and swabbing of built environment features among the Himba. Prof. Brooke Scelza (Anthropology, UCLA) has helmed a long-term demography and behavioral ecology research program among the community and recently launched a collaboration on breastmilk and breastfeeding among the Himba with Prof. Katie Hinde (Center for Evolution and Medicine, Arizona State University; Human Evolutionary Biology, Harvard), an anthropologist and co-advisor on my project. Thanks to these collaborators, I will be able to explore the importance of various environmental and cultural factors to rates of colonization by bifidobacteria
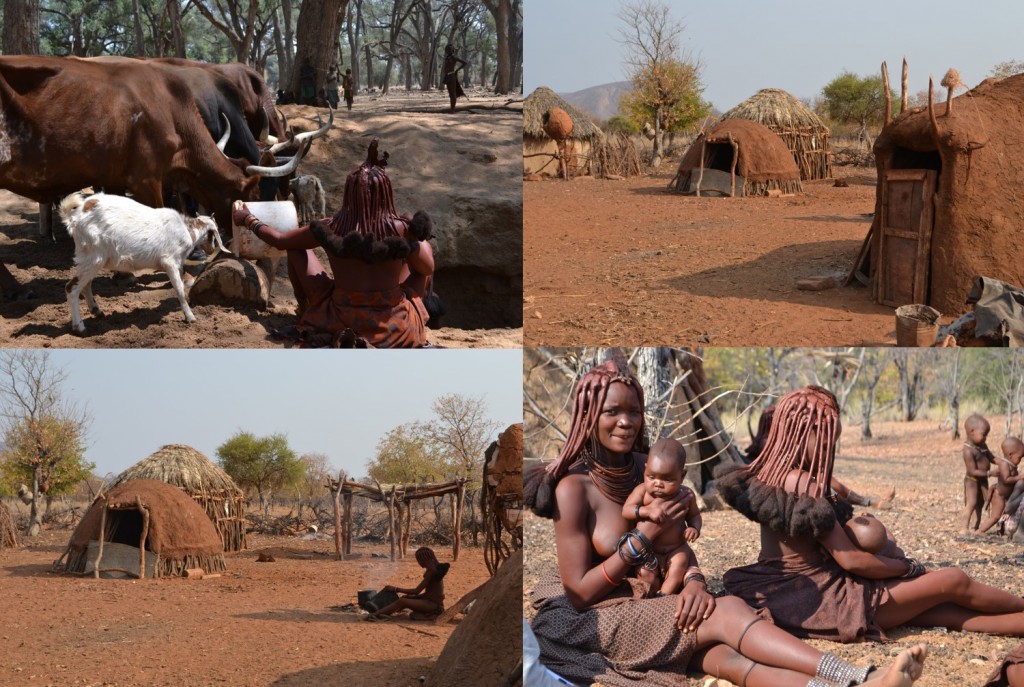

The past century has yielded unprecedented changes in both environmental and cultural factors that impact the human microbiota. It has become increasingly clear that these changes are often not without undesirable side effects. Lack of exposure to an inoculum of bifidobacteria suitable for the breast-fed infant gut was likely not an issue often faced by our ancestors. Our physiology, shaped by natural selection, may be adapted to “expect” exposure to bifidobacteria, whose presence provides crucial developmental cues and protections. The absence of these exposures is therefore of public health importance. Using populations in the Global South as a “more adaptive” point of reference, the available data suggest that the United States and other WEIRD populations may be facing widespread under-colonization by bifidobacteria. This project investigates the microbiology of the baby-associated built environment as a way to provide guidance on a potential mechanism that might be driving this phenomenon.
References
- Hinde, K. & Lewis, Z. T. Mother’s littlest helpers. Science (80-. ). 348, 1427—1428 (2015).
- Huda, M. N. et al. Stool Microbiota and Vaccine Responses of Infants. Pediatrics 134, e362—72 (2014).
- Lewis, Z. T. et al. Maternal Fucosyltransferase 2 Status Affects the Gut Bifidobacterial Communities of Breastfed Infants. Microbiome 3, 1—21 (2015).
- Henrich, J., Heine, S. J. & Norenzayan, A. The weirdest people in the world? Behav. Brain Sci. 33, 61—83; discussion 83—135 (2010).
- GrzeÅ›kowiak, Å. et al. Distinct gut microbiota in southeastern African and northern European infants. J. Pediatr. Gastroenterol. Nutr. 54, 812—6 (2012).
- Abrahamsson, T. R. et al. Low diversity of the gut microbiota in infants with atopic eczema. J. Allergy Clin. Immunol. 129, 434—40, 440.e1—2 (2012).
- Roos, S. et al. 454 pyrosequencing analysis on faecal samples from a randomized DBPC trial of colicky infants treated with Lactobacillus reuteri DSM 17938. PLoS One 8, e56710 (2013).
- Fouhy, F. et al. High-Throughput Sequencing Reveals the Incomplete, Short-Term Recovery of Infant Gut Microbiota following Parenteral Antibiotic Treatment with Ampicillin and Gentamicin. Antimicrob. Agents Chemother. 56, 5811—20 (2012).
- Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222—227 (2012).
- Marcobal, A. et al. Consumption of human milk oligosaccharides by gut-related microbes. J. Agric. Food Chem. 58, 5334—40 (2010).
- Yu, Z.-T., Chen, C. & Newburg, D. S. Utilization of major fucosylated and sialylated human milk oligosaccharides by isolated human gut microbes. Glycobiology 0, 1—12 (2013).
- Sela, D. A. et al. The genome sequence of Bifidobacterium longum subsp. infantis reveals adaptations for milk utilization within the infant microbiome. Proc. Natl. Acad. Sci. U. S. A. 105, 18964—9 (2008).
- Garrido, D., Kim, J. H., German, J. B., Raybould, H. E. & Mills, D. a. Oligosaccharide binding proteins from Bifidobacterium longum subsp. infantis reveal a preference for host glycans. PLoS One 6, e17315 (2011).
- Garrido, D., Dallas, D. C. & Mills, D. a. Consumption of human milk glycoconjugates by infant-associated bifidobacteria: mechanisms and implications. Microbiology 159, 649—64 (2013).
- Fukuda, S. et al. Bifidobacteria can protect from enteropathogenic infection through production of acetate. Nature 469, 543—7 (2011).
- Romond, M.-B. et al. Does the intestinal bifidobacterial colonisation affect bacterial translocation? Anaerobe 14, 43—8 (2008).
- Chichlowski, M., De Lartigue, G., German, J. B., Raybould, H. E. & Mills, D. a. Bifidobacteria isolated from infants and cultured on human milk oligosaccharides affect intestinal epithelial function. J. Pediatr. Gastroenterol. Nutr. 55, 321—7 (2012).
- Sheil, B. et al. Role of interleukin (IL-10) in probiotic-mediated immune modulation: an assessment in wild-type and IL-10 knock-out mice. Clin. Exp. Immunol. 144, 273—80 (2006).
- Tanabe, S., Kinuta, Y. & Saito, Y. Bifidobacterium infantis suppresses proinflammatory interleukin-17 production in murine splenocytes and dextran sodium sulfate-induced intestinal inflammation. Int. J. Mol. Med. 22, 181—185 (2008).
- Preising, J. et al. Selection of bifidobacteria based on adhesion and anti-inflammatory capacity in vitro for amelioration of murine colitis. Appl. Environ. Microbiol. 76, 3048—51 (2010).
- Roger, L. C., Costabile, A., Holland, D. T., Hoyles, L. & McCartney, A. L. Examination of faecal Bifidobacterium populations in breast- and formula-fed infants during the first 18 months of life. Microbiology 156, 3329—41 (2010).
- Roger, L. C. & McCartney, A. L. Longitudinal investigation of the faecal microbiota of healthy full-term infants using fluorescence in situ hybridization and denaturing gradient gel electrophoresis. Microbiology 156, 3317—28 (2010).
- Martin, M. A. & Sela, D. A. in Building Babies: Primate Development in in Proximate and Ultimate Perspective, Developments in Primatology: Progress and Prospects 37 (eds. Clancy, K. B. H., Hinde, K. & Rutherford, J. N.) 233—256 (Springer New York, 2013). doi:10.1007/978-1-4614-4060-4
- Young, S. L. et al. Bifidobacterial Species Differentially Affect Expression of Cell Surface Markers and Cytokines of Dendritic Cells Harvested from Cord Blood. Clin. Diagn. Lab. Immunol. 11, 686—690 (2004).
- De Wit, R. & Bouvier, T. ‘Everything is everywhere, but, the environment selects’; what did Baas Becking and Beijerinck really say? Environ. Microbiol. 8, 755—8 (2006).
- Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. U. S. A. 107, 11971—5 (2010).
- Chaban, B. & Seanhemmingsennrc-cnrcgcca, E. Characterization of the vaginal microbiota of healthy Canadian women through the menstrual cycle.
- Fanaro, S., Chierici, R., Guerrini, P. & Vigi, V. Intestinal microflora in early infancy: composition and development. Acta Paediatr. Suppl. 91, 48—55 (2003).
- Adlerberth, I. & Wold, a E. Establishment of the gut microbiota in Western infants. Acta Paediatr. 98, 229—38 (2009).
- Perez, P. F. et al. Bacterial imprinting of the neonatal immune system: lessons from maternal cells? Pediatrics 119, e724—32 (2007).
- Fernández, L. et al. The human milk microbiota: Origin and potential roles in health and disease. Pharmacol. Res. 69, 1—10 (2013).
- Urbaniak, C., Burton, J. P. & Reid, G. Breast, milk and microbes: a complex relationship that does not end with lactation. Womens. Health (Lond. Engl). 8, 385—98 (2012).
- Konya, T. et al. Associations between bacterial communities of house dust and infant gut. Environ. Res. 131, 25—30 (2014).