When we think of the built environment, we usually only include the indoor surfaces and air. A new study (preprint) posted last week on BioRxiv went through the roof, and looked at the microbes living on solar panels.
A highly diverse, desert-like microbial biocenosis on solar panels in a Mediterranean city – Pedro Dorado-Morales, Cristina Vilanova, Juli Pereto, Franscisco M Codoner, Daniel Ramon, Manuel Porcar – BioRxiv
The authors sampled solar panels on University of Valencia campuses during the summer solstice, both around noon (where temperatures are usually in the 90s) and at night. Analysis included culturing, 16S rRNA gene sequencing, metagenomics, and proteomics.
 Culturing yielded many bacteria with red, orange, or pink colors, that were very salt resistant. Ion Torrent sequencing of amplified ribosomal genes found hundreds of bacterial taxa with Deinococcus, Sphingomonas, Novosphingobium, and Hymenobacter as the dominant genera.
Culturing yielded many bacteria with red, orange, or pink colors, that were very salt resistant. Ion Torrent sequencing of amplified ribosomal genes found hundreds of bacterial taxa with Deinococcus, Sphingomonas, Novosphingobium, and Hymenobacter as the dominant genera.
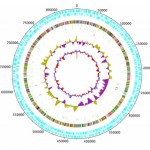 A Deinococcus “pangenome” could be constructed from the metagenomics sequences, with 2 chromosomes and 2 plasmids. Deinococcus spp. are often found at extreme environments such as deserts, but the pangenome from the solar panels appeared to contain a novel species. The paper did not delve into the genome much further, which would have been a good opportunity to find genes associated with living on a hot roof.
A Deinococcus “pangenome” could be constructed from the metagenomics sequences, with 2 chromosomes and 2 plasmids. Deinococcus spp. are often found at extreme environments such as deserts, but the pangenome from the solar panels appeared to contain a novel species. The paper did not delve into the genome much further, which would have been a good opportunity to find genes associated with living on a hot roof.
Metagenomic analysis showed that the solar panel microbiomes were similar to metagenomes from polar and saline desert environments.
In addition, the day and night samples showed differences in protein composition and signs of a metabolically active microbial community.
The manuscript is up on BioRxiv and open for comments.