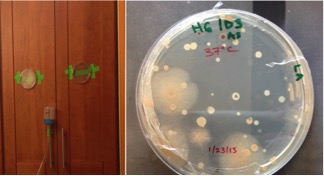
We just published an extended genome report from eight bacteria that were sequenced as part of the “Built Environment Reference Genome” (BERG) project. BERG is a microBEnet run effort to increase the number of reference genomes in the Built Environment (BE) by offering free genome sequencing to a number of labs working in the field. The goal of BERG was two-fold; 1) to get more genomes publicly available, and 2) to teach people – as much or as little as they wanted – about the process, and the power and utility of sequencing genomes.
These particular bacteria were isolated during aerosol sampling of bathrooms of residences in the San Francisco Bay area. We paired traditional cultivation techniques with particle sampling for sequence-based community composition analysis.
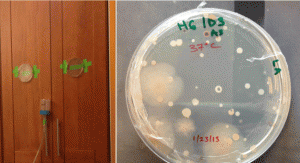
These specific eight isolates, spanning 5 genera, were sequenced because they are commonly identified in the built environment, originating from the dominant sources of indoor bacteria: the outdoor environment, human commensals, and premise plumbing. For instance, we included three species (four isolates) of staphylococci. Coagulase-negative staphylococci (CoNS) are typically benign inhabitants of the human skin and mucous membranes. Mycobacterium iranicum is a newly described species that has been isolated from all over the world and whose clinical significance is still under study. Pseudomonas oryzihabitans, despite its environmental origin, it has been recognized as a potential pathogen in recent years, especially in immunocompromised hosts. Other taxa include a Microbacterium and a Plantibacter.
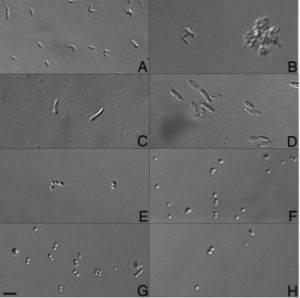
We explored the presence of genes associated with factors we thought to be relevant from the BE perspective. For example, the genome annotation of the CoNS revealed several genes associated with the ability to attach to a surface (e.g., putative adhesins), biofilm accumulation, and pro-inflammatory molecules with cytolytic and antimicrobial properties, among others. Additionally, we specifically looked for genes related to triclosan (TCS) resistance, a synthetic antimicrobial agent that is commonly used in a variety of home and personal care products. Although susceptibility to TCS or other antibiotics was not experimentally tested for our isolates, we found many genes associated with efflux pumps predicted to confer resistance to more than one class of antimicrobials (e.g., fluoroquinolones) or to specific antimicrobials (e.g., chloramphenicol). Gene components related to non-specific multidrug efflux pumps as induced by triclosan were found in almost all isolates, while P. oryzihabitans was the only isolate to contain an efflux pump predicted to offer triclosan resistance.
The chemical ecology of microorganisms on indoor surfaces is a component of BIMERC’s ongoing research efforts in the BE. We are studying the chemistry of biological interactions among microorganisms on residential indoor surfaces. The genomes of these eight isolates of bacteria will be valuable tools in our efforts to interpret future metagenomic and transcriptomic datasets, but also to understand these chemical interactions in vitro and to explore the basic microbiology of indoor microbes.