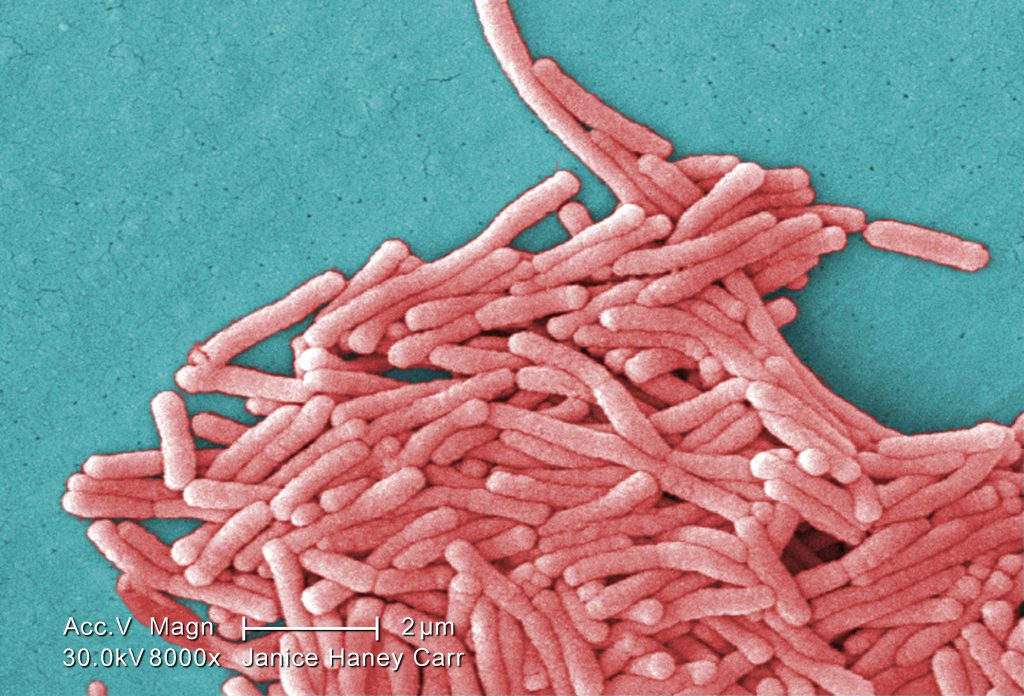
So this work is a spinoff of a big project that we were involved in (but almost all of the work was done by Amy Pruden’s lab at Virginia Tech). In the larger project, they examined the genomes of over 100 clinical isolates of Legionella pneumophila, as well as 10 clinical isolates from patients during the Flint Water Crisis. That work can be found here.
After genome sequencing it turned out that an additional 4 isolates were in fact not L. pneumophila but L. taurinensis, which had no sequenced genomes and is relatively poorly studied. The graduate student who lead this work, Connor Brown, noticed that this strain appeared to possess the genes for the RtxA Type 1 secretion systems (T1SS), a virulence factor thought to be unique to L. pneumophila. Further digging suggested RtxA, as well as other potentially novel T1SSs, are more widespread in this genus than previously reported. Our paper describing this work is entitled “Whole genome sequence analysis reveals the broad distribution of the RtxA type 1 secretion system and four novel putative type 1 secretion systems throughout the Legionella genus” and can be found here.
Abstract below:
Type 1 secretion systems (T1SSs) are broadly distributed among bacteria and translocate effectors with diverse function across the bacterial cell membrane. Legionella pneumophila, the species most commonly associated with Legionellosis, encodes a T1SS at the lssXYZABD locus which is responsible for the secretion of the virulence factor RtxA. Many investigations have failed to detect lssD, the gene encoding the membrane fusion protein of the RtxA T1SS, in non-pneumophila Legionella, which has led to the assumption that this system is a virulence factor exclusively possessed by L. pneumophila. Here we discovered RtxA and its associated T1SS in a novel Legionella taurinensis strain, leading us to question whether this system may be more widespread than previously thought. Through a bioinformatic analysis of publicly available data, we classified and determined the distribution of four T1SSs including the RtxA T1SS and four novel T1SSs among diverse Legionella spp. The ABC transporter of the novel Legionella T1SS Legionella repeat protein secretion system shares structural similarity to those of diverse T1SS families, including the alkaline protease T1SS in Pseudomonas aeruginosa. The Legionella bacteriocin (1–3) secretion systems T1SSs are novel putative bacteriocin transporting T1SSs as their ABC transporters include C-39 peptidase domains in their N-terminal regions, with LB2SS and LB3SS likely constituting a nitrile hydratase leader peptide transport T1SSs. The LB1SS is more closely related to the colicin V T1SS in Escherichia coli. Of 45 Legionella spp. whole genomes examined, 19 (42%) were determined to possess lssB and lssD homologs. Of these 19, only 7 (37%) are known pathogens. There was no difference in the proportions of disease associated and non-disease associated species that possessed the RtxA T1SS (p = 0.4), contrary to the current consensus regarding the RtxA T1SS. These results draw into question the nature of RtxA and its T1SS as a singular virulence factor. Future studies should investigate mechanistic explanations for the association of RtxA with virulence.