A moderately new paper is out that is an excellent example of how biases in DNA extraction can have major impacts on inferences from culture independent DNA studies. The paper is in what I think is a generally non open access journal (Computational and Structural Biotechnology Journal) for fortunately has been made open access
Source: Under-detection of endospore-forming Firmicutes in metagenomic data. Comput Struct Biotechnol J. 2015 Apr 25;13:299-306. doi: 10.1016/j.csbj.2015.04.002. eCollection 2015. Filippidou S, Junier T, Wunderlin T, Lo CC, Li PE, Chain PS, Junier P.
Abstract:
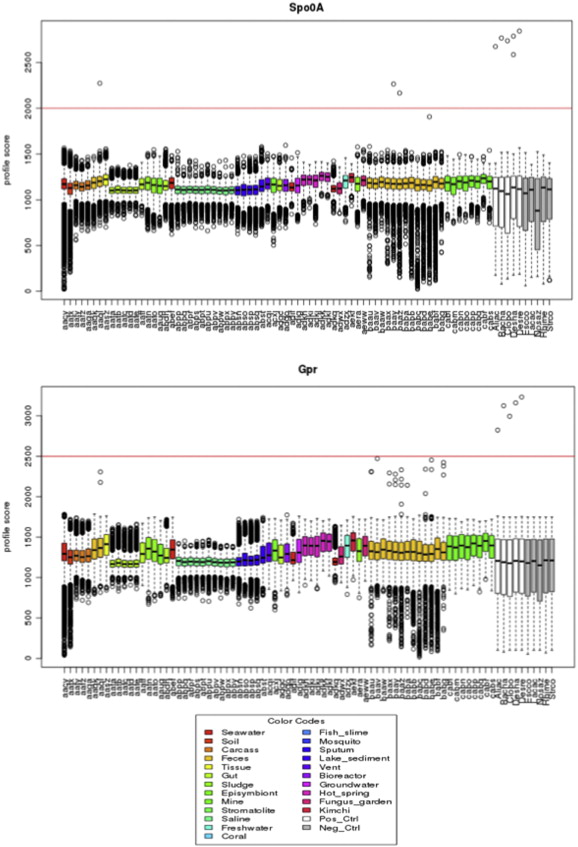
Microbial diversity studies based on metagenomic sequencing have greatly enhanced our knowledge of the microbial world. However, one caveat is the fact that not all microorganisms are equally well detected, questioning the universality of this approach. Firmicutes are known to be a dominant bacterial group. Several Firmicutes species are endospore formers and this property makes them hardy in potentially harsh conditions, and thus likely to be present in a wide variety of environments, even as residents and not functional players. While metagenomic libraries can be expected to contain endospore formers, endospores are known to be resilient to many traditional methods of DNA isolation and thus potentially undetectable. In this study we evaluated the representation of endospore-forming Firmicutes in 73 published metagenomic datasets using two molecular markers unique to this bacterial group (spo0A and gpr). Both markers were notably absent in well-known habitats of Firmicutes such as soil, with spo0A found only in three mammalian gut microbiomes. A tailored DNA extraction method resulted in the detection of a large diversity of endospore-formers in amplicon sequencing of the 16S rRNA and spo0A genes. However, shotgun classification was still poor with only a minor fraction of the community assigned to Firmicutes. Thus, removing a specific bias in a molecular workflow improves detection in amplicon sequencing, but it was insufficient to overcome the limitations for detecting endospore-forming Firmicutes in whole-genome metagenomics. In conclusion, this study highlights the importance of understanding the specific methodological biases that can contribute to improve the universality of metagenomic approaches.
I noticed this paper a few months ago and posted about it to Twitter:
Seems important for microbial ecology:Under-detection of endospore-forming Firmicutes in metagenomic data http://t.co/ElscGyijB8 #microBEnet
– Jonathan Eisen (@phylogenomics) May 2, 2015
But only now getting around to blogging about it. This is definitely worth a look.
I made a Storify of some of the responses (jsut a few) on Twitter. I am going to try to embed it here in the comment