So – just a quick post for now. More details to come. But I wanted to get a little bit about this out there.
For a few years I have been teaching a mid-level course at UC Davis on “DNA sequencing based studies of microbial diversity’. The course has evolved over the years from a primarily lecture based course into a course focused much more on reading scientific papers and on how to read such papers. I taught the course this winter and in collaboration with my PhD student Cassie Ettinger who served and the TA for the course, we went even more extreme in terms of the scientific reading part. The entire course was organized around a single paper. We had the students read the paper at the beginning of the course with the full knowledge that they would not understand a lot of it. And then for the next 10 weeks we went through many topics relating to DNA sequencing and microbes organized in a large part around understanding the various parts of this one paper. I really liked how this turned out and I plan to definitely do this in the future for other courses.
For today I am going to provide an overview of the course and then over the next few weeks I will try to post additional details.
The main paper.
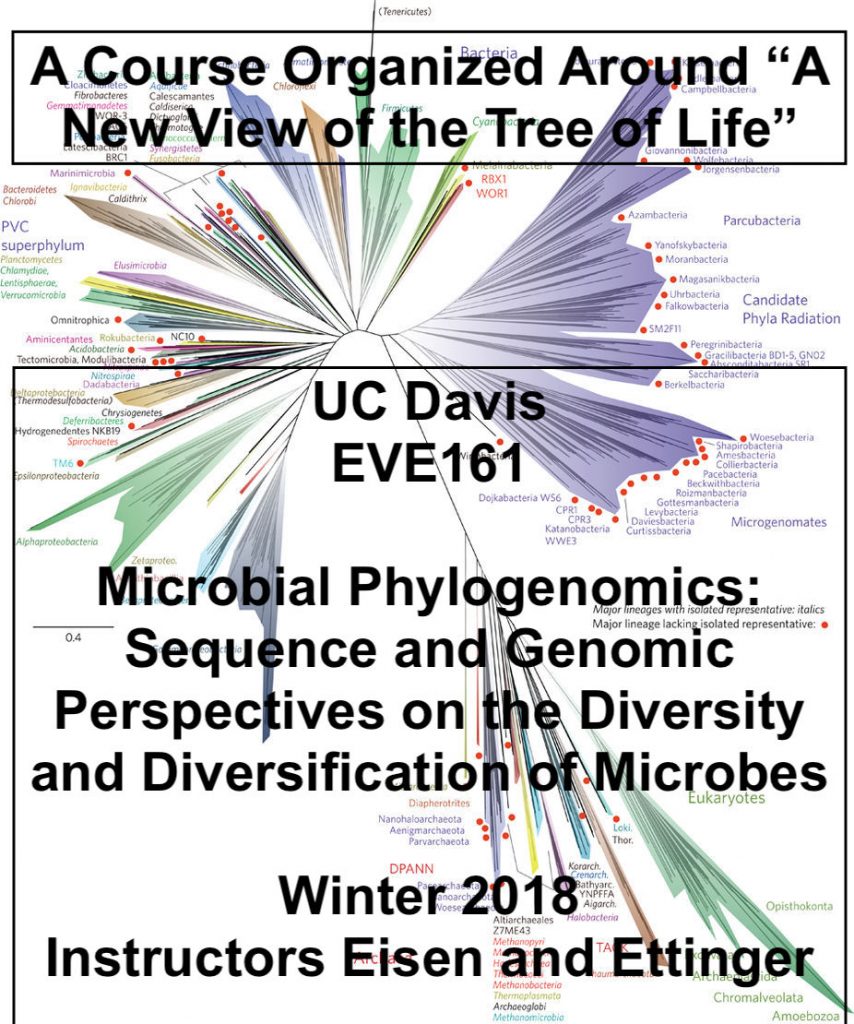
The paper we organized the course around was Hug et al. “A New View of the Tree of Life.”
Full citation: Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, Butterfield CN, Hernsdorf AW, Amano Y, Ise K, Suzuki Y, Dudek N, Relman DA, Finstad KM, Amundson R, Thomas BC, Banfield JF. 2016. A new view of the tree of life. Nature Microbiology 2016 1:16048. doi: 10.1038/nmicrobiol.2016.48.
We chose this paper for a few reasons including
- I really like it
- It covers directly or indirectly all six of the main topics I normally cover in this class: sequencing, rRNA sequencing of cultured organism, culture independent rRNA studies, genome sequencing of cultured organisms, metagenomics, and phylogeny.
- It is open access
- It is pretty new
- I have used it in classes previously (but only for single day topics) and thus was at least familiar with it
The Course Description
For the course, the basic plan was to use this paper as an organizing principle both for the scientific topics of the course and also in relation to the general topic of “how to read a scientific paper”. However, we did not fully reveal this at the beginning. Instead I kind of kept the course description as I have done before – really general. For example, here is the syllabus we posted to the UC Davis Canvas Site
EVE161
Microbial Phylogenomics
Sequence and Genomic Perspectives on the Diversity and Diversification of Microbes
Winter 2018
3 UnitsCourse Description
The course focuses on the use of DNA sequencing in studies of the diversity and the diversification of microorganisms. Topics covered include both methods (e.g., DNA sequencing, PCR, phylogenetics, genome sequencing, phylogenomics, metagenomics), the diversity of microbes (e.g., the tree of life, microbial communities and microbiomes, mechanisms of diversification such as lateral gene transfer). The course also has a major emphasis on how to read the primary scientific literature.
Course Objectives
After taking this course students should have
- A better understanding of the history of sequence based studies of microbial diversity and current practice in sequence based studies of microbial diversity,
- A broad view of what we know about microbial diversity and
- Improved ability to read and analyze a research paper.
Evaluation
Grades will be determined by the following components:
-
Attendance and class participation 10 %
-
Daily assignments 30 %
-
Midterm 20 %
-
Research project (class presentation and write up) 20%
-
Final exam 20%
And then prior to the first class we sent an announcement to all the students
Reading for EVE161 1st class on 1/9
Jan 4 at 10:50am
No unread replies.No replies.
- Class 15: Era IV – Metagenomics Large insert
- Class 16: Era IV – Metagenomics Shotgun metagenomics
- Class 17: Era IV – Metagenomic Predictions
Return to the beginning
- Class 18: Hug et al.
- Class 19: Hug et al.
In the next few weeks I will post more details about the course, the papers we read, the assignments we gave, and other information.