So this was a very interesting paper to be involved in. On the one hand all our lab really did was to facilitate some genome sequencing and help with the assemblies, validation of those assemblies, and some genomics work. Standard fare for the lab. But… these were over 100 Legionella pneumophila isolates collected from either patients or environmental sources associated with the Flint Michigan outbreak where a dozen people died and a myriad of lawsuits are presumably still pending. All of the substantive analyses and writing were done by our collaborators at Virginia Tech, but the experience of even being tangentially involved in a project on the edges of so much media scrutiny was new territory.
Here’s the Abstract of the paper:
During the water crisis in Flint, Michigan, USA (2014–2015), 2 outbreaks of Legionnaires’ disease occurred in Genesee County, Michigan. We compared whole-genome sequences of 10 clinical Legionella pneumophila isolates submitted to a laboratory in Genesee County during the second outbreak with 103 water isolates collected the following year. We documented a genetically diverse range of L. pneumophila strains across clinical and water isolates. Isolates belonging to 1 clade (3 clinical isolates, 3 water isolates from a Flint hospital, 1 water isolate from a Flint residence, and the reference Paris strain) had a high degree of similarity (2–1,062 single-nucleotide polymorphisms), all L. pneumophila sequence type 1, serogroup 1. Serogroup 6 isolates belonging to sequence type 2518 were widespread in Flint hospital water samples but bore no resemblance to available clinical isolates. L. pneumophila strains in Flint tap water after the outbreaks were diverse and similar to some disease-causing strains.
So scientifically, the major take home message was that there was a surprisingly amount of diversity in both environmental and clinical Legionella strains found after the outbreak. One could easily imagine a scenario where a single strain is responsible for an outbreak but that doesn’t appear to have been the case here.
In terms of “what does it mean?” I was struck by a couple of major limitations in this whole kind of study. Firstly, water sources aren’t routinely monitored for Legionella (everywhere, not just in Flint) so there are no water samples from during the actual outbreak which would obviously be useful. Secondly, they were only able to obtain 10 clinical isolates even though ~90 people were infected during the outbreak. That’s because the bacteria is not often cultured from sick patients. Much better environmental monitoring, combined with better collection of patient isolates would presumably allow source tracking in a future outbreak which could lead to improved outcomes and prevention.
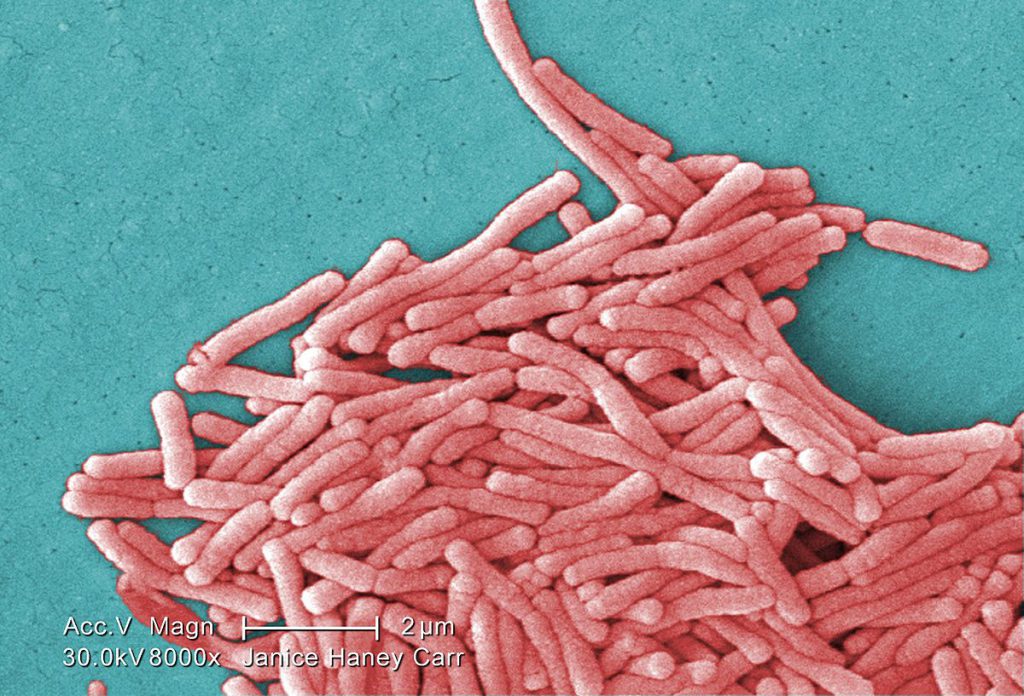
One word and two phrases come to mind regarding. the Flint LD outbreaks:
1) Indigenous
2) “DEAD-leg” scenario
3) Initial-Precipitating-Event (IPE)