This article (“Collection of SARS-CoV-2 Virus from the Air of a Clinic within a University Student Health Care Center and Analyses of the Viral Genomic Sequence”) caught my attention initially because of the air sampling aspect, but upon reading the Abstract I was struck by something else. Here they did air sampling in a …
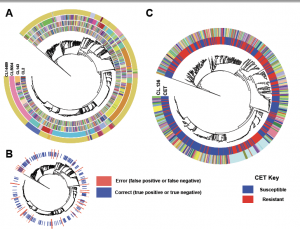
This preprint seems like it may be of interest to folks: Precise prediction of antibiotic resistance in Escherichia coli from full genome sequences | bioRxiv Basically, the authors showed that, using a machine learning approach, they can quite accurately predict antibiotic resistance in E. coli from whole genome data. The emergence of microbial antibiotic resistance …
(Note: This post was updated on 3/26/18 after re-running some of our analysis based on suggestions in the comments and on Twitter). We were very excited to get our MinION sequencer from Nanopore 1.5 years ago… so many potential applications; sequencing in real time, use in the classroom, use in the field, not having to …
Over the last several years our lab has spent a fair bit of effort on collecting and sequencing genomes of bacteria from the built environment. Genome sequences are useful for a number of reasons including facilitating metagenomics and understanding the actual metabolic capacity of particular microbes. In particular we hope that genome sequences will facilitate …
Isolating interesting bacteria and sequencing their genomes has never been easier and our lab has worked extensively on this topic, particularly in the context of undergraduate education. Our first foray into this field came in 2011 with a group of students working on the “Built Environment Genomes Project” and eventually culminated in our Swabs to …
You can check out David Coil’s introduction to the project here. The workflow pre-print is hosted by Peer J here. Feel free to check it out! We would love any comments or suggestions. I was first introduced to the swabs to genome workflow project a little over a year ago. I had just started in the …
Almost exactly a year ago we finally wrapped up our undergraduate project focusing on sequencing and assembling reference genomes from the built environment. This project aimed to take undergrads through every step from starting with a swab to ending with a published genome announcement and data in NCBI. Over the course of the work, we …
With the publication of the 6th and last genome paper to come out of our Undergraduate Genome Sequencing Project I thought this would be a good time to reflect on how it all went. To summarize, we had a group of undergraduate students go out into the built environment and attempt to find microbes whose …