So – we have been having a running discussion with people in my lab about one key issue in microbiome studies – how does one store samples prior to doing DNA extractions and does it matter? As background for those who do not do this kind of work – the general principle behind DNA based …
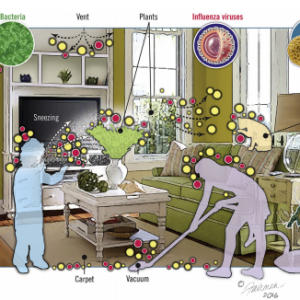
By Amanda Makowiecki 1st Year Mechanical Engineering PhD Student Miller Research Group, University of Colorado Boulder Researchers at the University of Oregon recently published a paper examining the connection between architectural design and microbial diversity in our buildings (Kembel et al. 2014). Although occupancy type was identified as the strongest predictor of microbial variation, several …
NOTE (3-12-15): THESE RESULTS ARE INCORRECT. We have left them here for comparison. A blog post describing the problem is here and the correct information can be found here. We’ve finished analyzing all the data from the “Microbial Playoffs” part of Project MERCCURI (described here). Each microbe that was chosen to fly to the International …
IBM and Mars put out a joint press release today announcing a new effort to use metagenomics to study microorganisms in the food supply chain. The new initiative, called the Sequencing the Food Supply Chain Consortium (SFSC), will use metagenomics and metatranscriptomics to establish what they call a “microbial baseline” that can later be used …
From the perspective of biothreat detection, one of the biggest questions is “what is the background?”. If you sample the air near a farm, there’s a good bet that you’ll find anthrax (or at least a close relative indistinguishable by a 16S PCR survey). But that’s just because anthrax lives in the soil, not because …
Saw an interesting talk yesterday from Karen Guillemin of the University of Oregon. I made a storification of the Twitter posts from the talk. See it at the end of this post. I note – I think there are some important lessons in the work on germ free animals that could be applied to studies …
Compiling some of the more interesting tools I have seen recently. Some I have plyed with but most I have just looked at the papers briefly. Microbiome | Abstract | VizBin – an application for reference-independent visualization and human-augmented binning of metagenomic data. Global biogeographic sampling of bacterial secondary metabolism GrammR: Graphical Representation and Modeling …
There is an abundance of literature on how microbes can obtain antibiotic resistance, but not as much about how antibiotic resistance can spread. Jonathan drew my attention to this article today, which highlights the fact that antibiotic resistance can be spread through the air. While I didn’t find the conclusions all that surprising, I was …
I was recently contacted by a SRA curator to submit the raw pacbio datasets that go with genomes that were deposited to NCBI. I did go through with the submission and will share what I did and my experience doing so. Be prepared to answer a lot of questions regarding your project as well as …
Crosspsting from my Tree of Life blog This is so cool: Tangible Interactive Microbiology for Informal Science Education. Abstract: We present an interactive platform that enables human users to interface with microbiological living cells through a touch-screen, thereby generating a tangible interactive experience with the microscopic world that is hidden to most people. Euglena gracilis, single-celled …