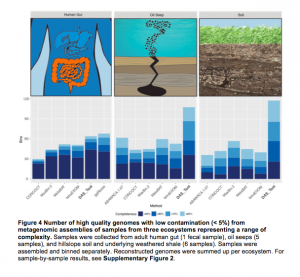
Saw an interesting Tweet check out this novel metagenomics tool to reconstruct thousands of high-quality genomes @ChrisMKSieber #banfieldlab https://t.co/67bk8CbFmP — Alexander Probst (@AlexJProbst) February 13, 2017 And this tool may be of interest – it is from a new preprint in BioRXiv. See abstract below: Microbial communities are critical to ecosystem function. A key objective …
There are a number of cases where determining the relationship between microbes is at the center of a research question. Are the microbes inhabiting a building the same as those inhabiting its tenants? Are the microbes in a hospital room the same as those that colonize newborn babies? Is the E. coli living on a …
There is an increasing number of studies with a large number genomes recovered from isolate, metagenome, or single cell sequencing. To bridge the gap between the available genome sequences and available phenotype information, we have developed Traitar, a bioinformatics software to phenotype bacteria based on their genome sequence (see workflow below) . Traitar includes phenotype models for …
There has been some interest in our recent preprint describing Oxford Nanopore MinIONTM sequencing for 16S rRNA microbiome characterization and I was asked to write a post for microbenet on this technology. Disclaimers – this paper is a work in progress – our paper has not yet been peer-reviewed and we are continuing to revise …
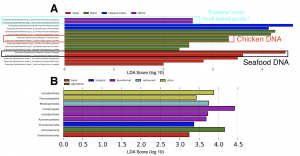
Amidst the November/December holiday chaos, myself and co-authors were proud to witness the publication of a neat new paper focused on ATM keypads in New York City. Yes, just like all other surfaces in the Built Environment, those ATM keypads are harboring lots of microbes and bits of orphaned DNA! This ATM keypad study was work that …
Just got back from the Pacific Symposium in Biocomputing. The meeting had some aspects that may be of interest to various folks. Electronic proceedings of the meeting are here. I talked in a session on functional predictions. I was asked to do this at the last minute so I made my slides by hand. Here …
Kevin Van Den Wymelenberg and Jessica Green, of the Biology and the Built Environment Center (BioBE), are currently seeking a microbial ecology Research Associate / Research Assistant Professor / Research Associate Professor (non-tenure track faculty) to investigate fundamental questions surrounding the role of microorganisms (bacteria, archaea, fungi, protists, and viruses) in the built environment and …
The Department of Statistics at Oregon State University is hosting a summer REU (Research Experiences for Undergraduates) program that focuses on microbiome data analysis. This is a unique, paid research and training opportunity for STEM undergraduate students who want an interdisciplinary research experience in both biology and statistics. During a ten week training period, students will be …
I was at a meeting a few weeks ago where the topic of fungal diversity surveys was discussed and many people there commented on how ITS based surveys (one of the main approaches for culture independent studies of fungi) had some limitations. ITS stands for the “internal transcribed sequence” and it is a region in …
This is definitely worth checking out. Titus Brown and Harriet Alexander taught a two day metagenomics workshop at UCSD and they have written up some details about it and posted videos. Wrap up blog: titus-blog/2016-metagenom ics-workshop-at-sio.rst at public · ctb/titus-blog · GitHub All the docs here: http://2016-metagenomics-sio.readthedocs.io/en/latest/ Videos