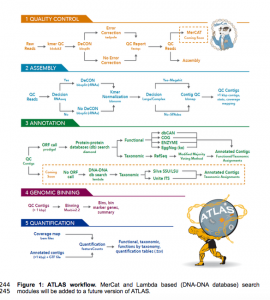
OK I have not tried this out yet but this could be of interest. ATLAS (Automatic Tool for Local Assembly Structures) is a comprehensive multi-omics data analysis pipeline that is massively parallel and scalable. ATLAS contains a modular analysis pipeline for assembly, annotation, quantification and genome binning of metagenomics and metatranscriptomics data and a framework …
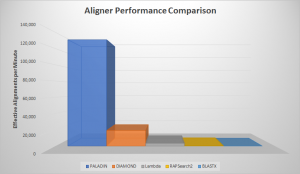
With our paper recently accepted for publication and many months of practical application, we’re excited to spread the word about our new tool for characterizing metagenomes, PALADIN! Short for Protein ALignment And Detection INterface, PALADIN was our response to the inherent problem of limited references in shotgun metagenomic sequencing. While DNA barcoding involving 16S, 18S, …
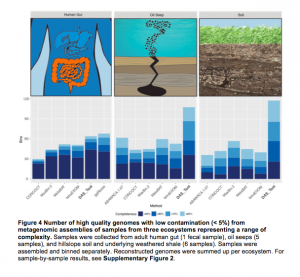
Saw an interesting Tweet check out this novel metagenomics tool to reconstruct thousands of high-quality genomes @ChrisMKSieber #banfieldlab https://t.co/67bk8CbFmP — Alexander Probst (@AlexJProbst) February 13, 2017 And this tool may be of interest – it is from a new preprint in BioRXiv. See abstract below: Microbial communities are critical to ecosystem function. A key objective …
There is an increasing number of studies with a large number genomes recovered from isolate, metagenome, or single cell sequencing. To bridge the gap between the available genome sequences and available phenotype information, we have developed Traitar, a bioinformatics software to phenotype bacteria based on their genome sequence (see workflow below) . Traitar includes phenotype models for …
Just a quick post here. Studies of the microbiology of built environments that house animals (e.g., aquaria, farms, animal shelters, zoos, etc) are of growing interest for multiple reasons. In that regard this paper might be of interest to some – because it covers some topics that are sometimes neglected in this general area – viral diversity …
This is definitely worth checking out. Titus Brown and Harriet Alexander taught a two day metagenomics workshop at UCSD and they have written up some details about it and posted videos. Wrap up blog: titus-blog/2016-metagenom ics-workshop-at-sio.rst at public · ctb/titus-blog · GitHub All the docs here: http://2016-metagenomics-sio.readthedocs.io/en/latest/ Videos
In January 2014 I wrote this post about “microbiome” sequencing services. I have gotten many outside requests for the following information — what places (companies, Universities, government agencies, etc) provide contract services for rRNA PCR and sequencing? Source: Request — Information on Places that do rRNA sequencing as a service — microBEnet: the microbiology of the …
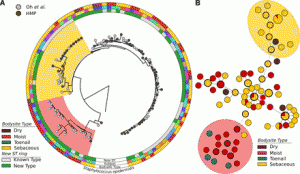
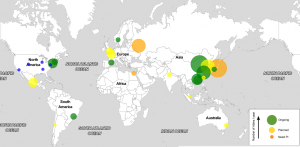
New software/tool of possible interest: Source: MetaMLST: multi-locus strain-level bacterial typing from metagenomic samples Software: http://segatalab.cibio.unitn.it/tools/metamlst/ Abstract: Metagenomic characterization of microbial communities has the potential to become a tool to identify pathogens in human samples. However, software tools able to extract strain-level typing information from metagenomic data are needed. Low-throughput molecular typing schema such as Multilocus …
Videos of talks from the recent DOE-JGI User Meeting have been posted. There is also a Storify wrap up of the meeting: [<a href=”//storify.com/doe_jgi/2016-doe-jgi-genomics-of-energy-environment-meetin” target=”_blank”>View the story “2016 DOE JGI Genomics of Energy & Environment Meeting” on Storify]
One afternoon while making my way through another DNA extraction, I was eavesdropping over the lab benches, per usual, trying to keep my mind occupied. I overheard my project manager and another undergraduate discussing an unusual project. “Do you want to contact the Sacramento authorities to get clearance to swab the light rail, because talking …