So – just a quick post for now. More details to come. But I wanted to get a little bit about this out there. For a few years I have been teaching a mid-level course at UC Davis on “DNA sequencing based studies of microbial diversity’. The course has evolved over the years from a …
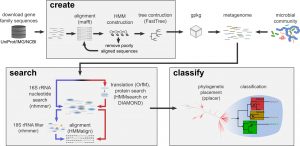
This could be of use to many people: Source: GraftM: a tool for scalable, phylogenetically informed classification of genes within metagenomes | Nucleic Acids Research | Oxford Academic Large-scale metagenomic datasets enable the recovery of hundreds of population genomes from environmental samples. However, these genomes do not typically represent the full diversity of complex microbial …
Just a quick post here. There is a paper of interest I thought I would call attention to: Source: SILVA, RDP, Greengenes, NCBI and OTT — how do these taxonomies compare? | BMC Genomics by Monika Balvočiūtė and Daniel H. Huson Abstract: Background A key step in microbiome sequencing analysis is read assignment to taxonomic units. …
So – I am teaching in BIS002C – Biodiversity and the Tree of Life again at UC Davis. And I thought it might be of interest to share (and get feedback on) some of what I am teaching for this course. Quick summary – this is the third course in an Intro Bio series …
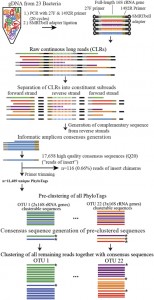
This seems like it could be of interest to microbial community researchers: The ISME Journal – High-resolution phylogenetic microbial community profiling Abstract (with bolding by me) Over the past decade, high-throughput short-read 16S rRNA gene amplicon sequencing has eclipsed clone-dependent long-read Sanger sequencing for microbial community profiling. The transition to new technologies has provided more …