Latest Other Posts
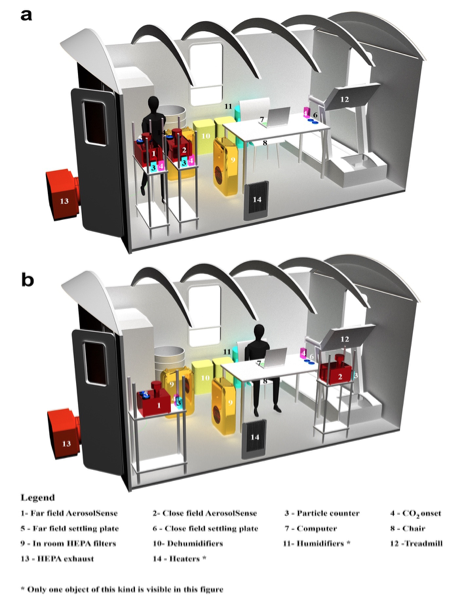
(posting on behalf of Dr. Karen Dannemiller) The Indoor Air 2024 conference from the International Society for Indoor Air Quality and Climate (ISAIQ) will take place in beautiful Honolulu, HI. Have your research highlighted as part of Indoor Air 2024’s Conference Countdown Challenge! Starting July 10th, 2023, submissions of images related to the conference themes ...
 Job posting: Tenure-Track Faculty Position in Environmental Microbiology at Virginia Tech
Job posting: Tenure-Track Faculty Position in Environmental Microbiology at Virginia Tech
Job Posting: The Charles E. Via, Jr. Department of Civil and Environmental Engineering (CEE) at Virginia Tech invites applications for a tenure-track faculty position in environmental engineering at the rank of Assistant Professor. We are seeking candidates motivated to contribute to a collegial, interdisciplinary community in several core mission areas of the university. The ...
A new viral disease surveillance tool: Indoor dust as a matrix for surveillance of COVID-19, published in mSystems. Throughout the COVID-19 pandemic, large scale monitoring of SARS-CoV-2 has helped to prevent spread and inform public health decisions. This has largely been done through individual sampling (ie nasal and saliva sample collection) and through environmental surveillance ...
Search MicroBE.net
Support microBEnet
The microBEnet website is a hub for information about the microbiology of the built environment. We strive to be as up-to-date as possible on new papers, developing technologies, upcoming events, etc. Originally and currently funded by the Alfred P. Sloan Foundation, ongoing donations help support the website and the addition of new content.
Upcoming Events
There are no upcoming events at this time.
- Microbial partners of crop plant agro-biodiversity December 4, 2018
- Never Home Alone: August 20, 2018
- Fermented Foods: Harnessing the Power of Microorganisms April 6, 2018